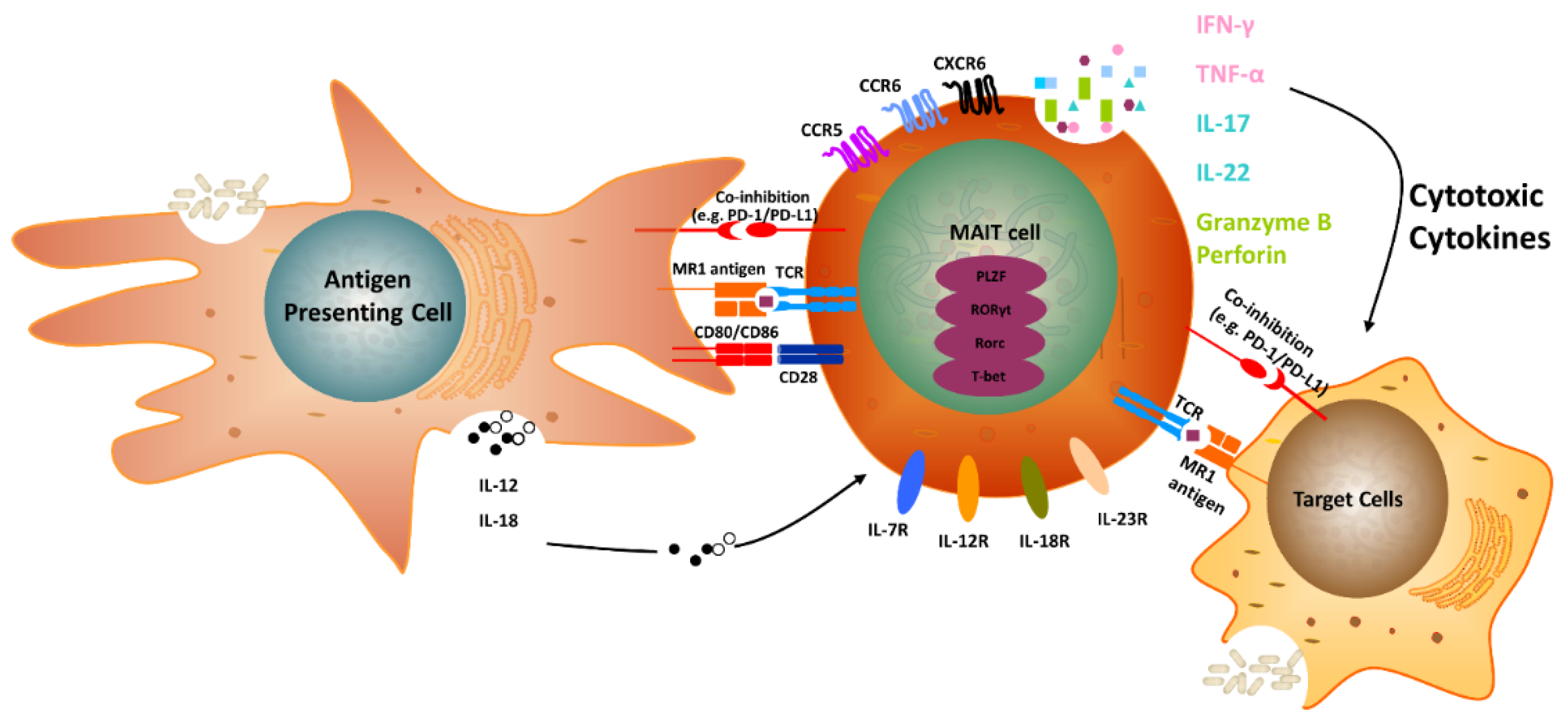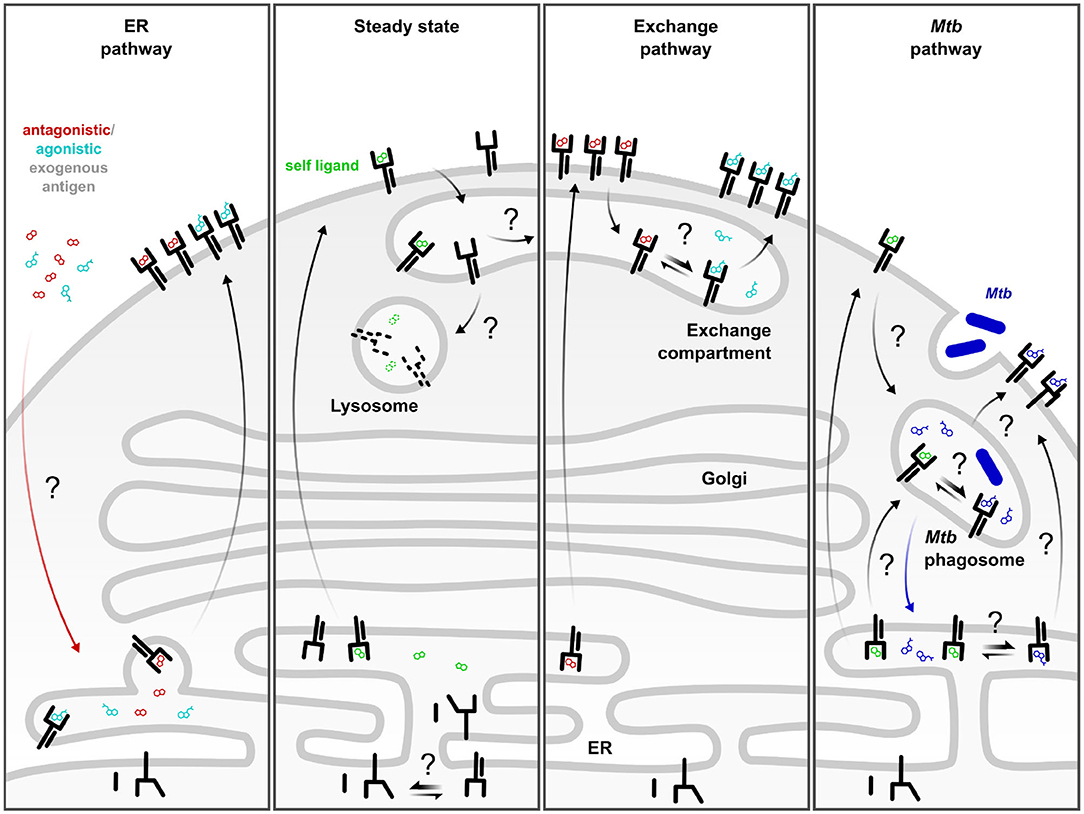
Frontiers | Covering All the Bases: Complementary MR1 Antigen Presentation Pathways Sample Diverse Antigens and Intracellular Compartments
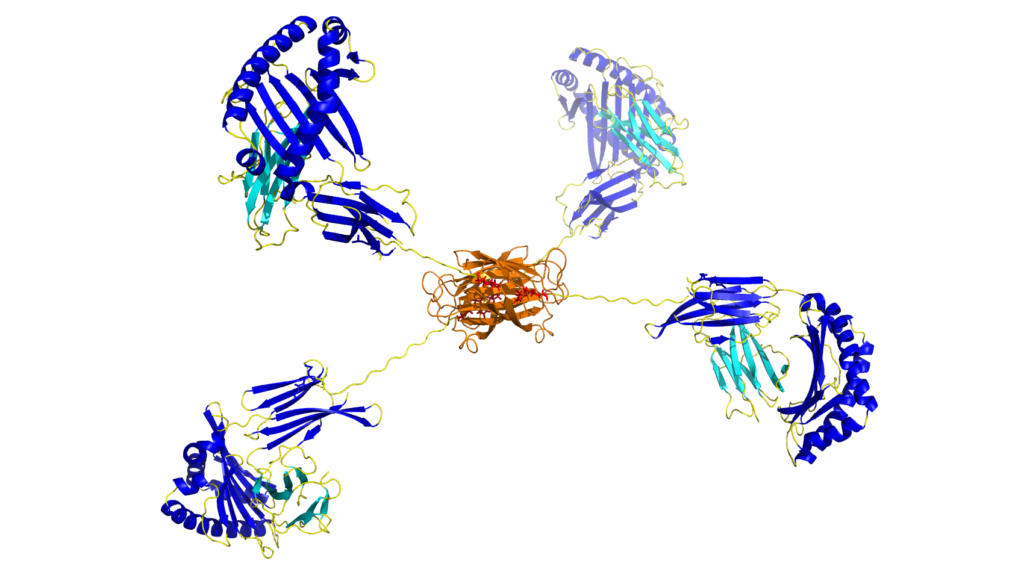
MR1 tetramers - ProImmune - Mastering Immunity _ MHC pentamers, CD1d tetramers, custom peptide synthesis, immunoassays, T cell epitope discovery, HLA tissue typing, cellular analysis, ligand binding assays, immune response

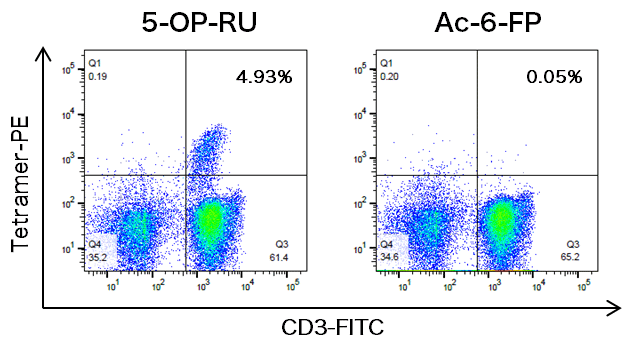
Figure 3 | Generation of MR1-Restricted T Cell Clones by Limiting Dilution Cloning of MR1 Tetramer+ Cells | SpringerLink
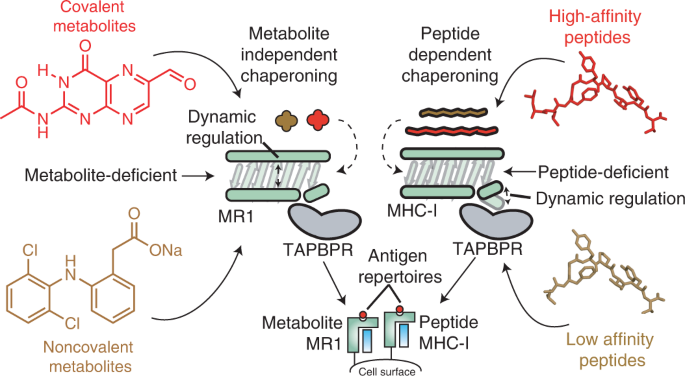
TAPBPR employs a ligand-independent docking mechanism to chaperone MR1 molecules | Nature Chemical Biology
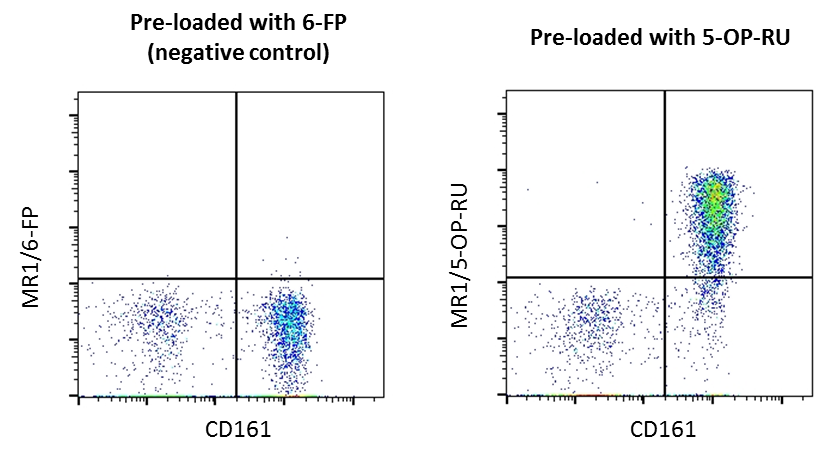
MR1 tetramers - ProImmune - Mastering Immunity _ MHC pentamers, CD1d tetramers, custom peptide synthesis, immunoassays, T cell epitope discovery, HLA tissue typing, cellular analysis, ligand binding assays, immune response

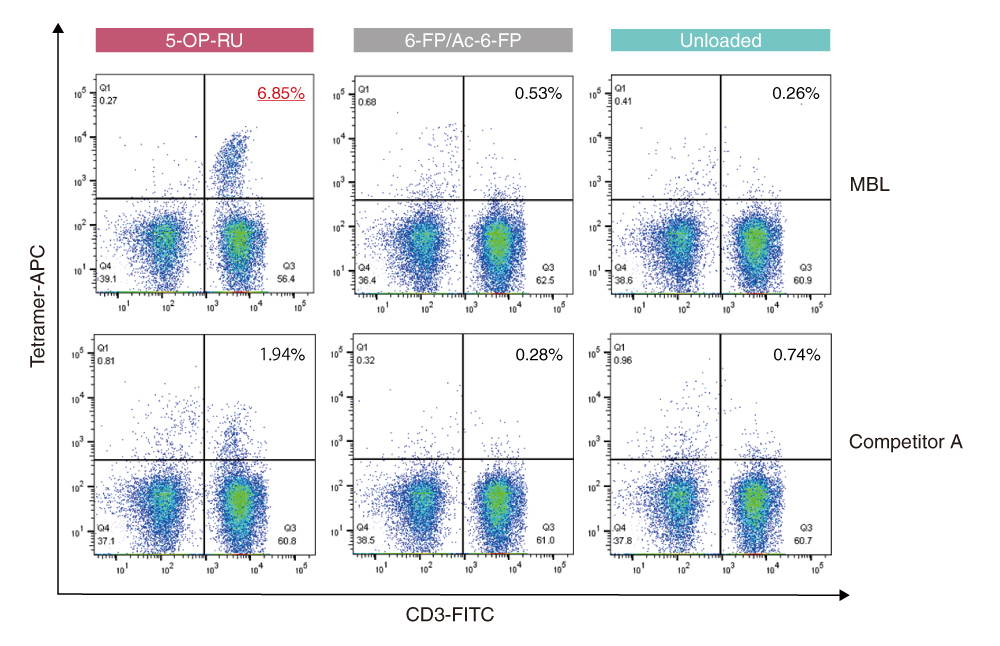
MR1-Restricted T Cells with MAIT-like Characteristics Are Functionally Conserved in the Pteropid Bat Pteropus alecto - ScienceDirect

Linkage map construction on based of F 2 population derived from MR-1... | Download Scientific Diagram

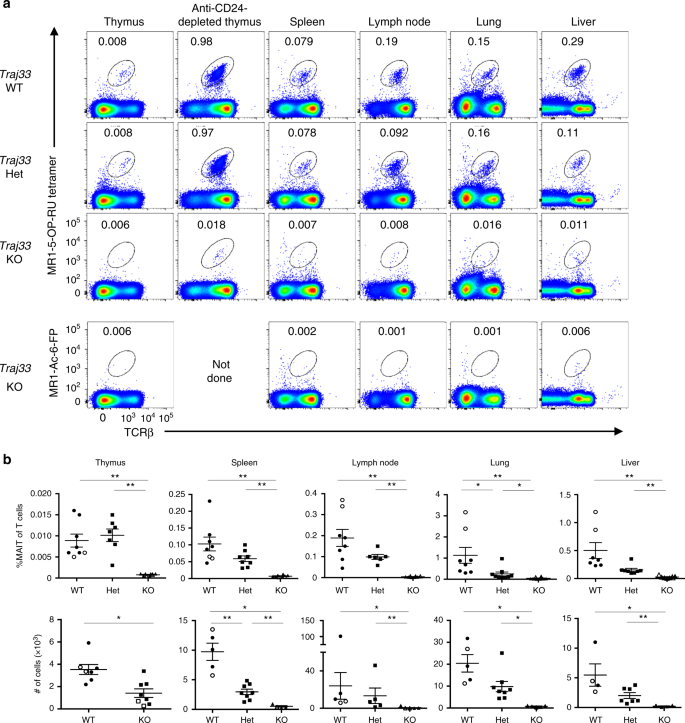
Bacteria-dependent population expansion of MAIT cells, activation at... | Download Scientific Diagram

How MR1 Presents a Pathogen Metabolic Signature to Mucosal-Associated Invariant T (MAIT) Cells: Trends in Immunology